
Concept explainers
To review:
The structures ofproducts of the reaction between nitrogenase complex and substrates hydrogen cyanide, dinitroge, nnd acetylene.
Introduction:
Nitrogenase complex is present in all the species that can fix nitrogen. It is produced by bacteria such as cyanobacteria. This reduces nitrogen (N2) to ammonia (NH3). This is the only enzyme that can catalyze the reaction for nitrogen fixation.
Explanation of Solution
The nitrogenase complex consists of two proteins called dinitrogenase as well asdinitrogenase reductase. The dinitrogenase is heterotetramer of MoFe protein. The dinitrogenase reductase is called Fe(ferrous) protein. It contains Mg-ATP (magnesium-adenosine triphosphate)-binding sites. In the reaction catalyzed by this complex, there is a transfer of electrons from (4Fe-4S) cluster in Fe protein. The Fe transfers the electrons to Mo-Fe protein. The transfer of electrons leads to a reduction of H+. The incoming N2 exchanges withH+ within the active site to form stable intermediates. So, the reaction of nitrogenasecomplex with the given substrates is as follows:
When nitrogenase complex reacts with cyanide, it gives two product, sethylamine and methan, elong with ammonia:
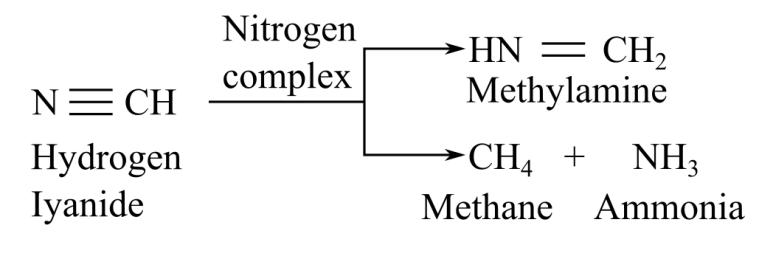
When nitrogenase complex reacts with acetylene, it gives ethylene as a product:
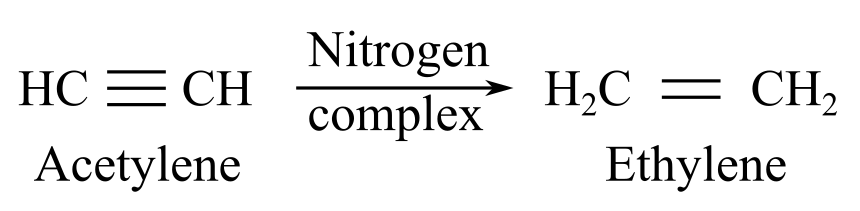
When nitrogenase complex reacts with dinitrogen, it gives hydrazine or ammonia as a product:
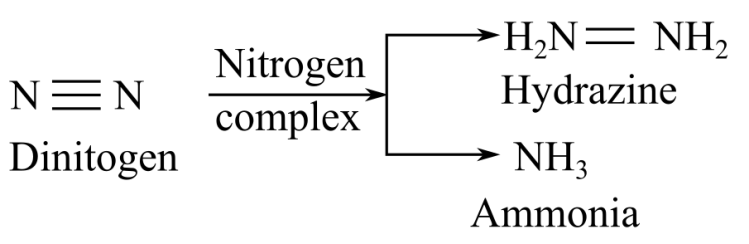
Thus, it can be concluded that the products of substrates hydrogen cyanide, acetylen, endnitrogen are methylamine, ethylene, and hydrazine, respectively.
Want to see more full solutions like this?
Chapter 14 Solutions
Biochemistry: The Molecular Basis of Life
- (b) The values of kinetic parameters for a variety of synthetic ester and peptide substrates ofa- chymotrypsin are compared for the reaction scheme below where ES is the Michaelis complex, ES' E+S Substrate K₁ N-Ac-Trp–OC₂H5 N-Ac-Phe-OC2H5 N-Ac-Leu-OC2H5 N-Ac-Phe-CONH2 N-Ac-Tyr-p-nitro-anilide ES K₂ 3.5 13.0 3.2 1.7 K-₁ is the acylenzyme, P₁ is the first product to be released, P2 is the second product, k2 is the acylation rate constant, k3 is the deacylation rate constant, and kcat = k2 k3/( k2 + k³). In the table below, N- Ac = N-acetyl; -CONH₂ = carboxamide; -OC₂H5 = ethyl ester; p-nitroanilide = −NH-C6H4-NO2 simulating a peptide group. ES' K2 (S-¹) K3 (S-1) 0.84 2.2 0.19 P₁ K3 0.073 H → E+ P2 Kcat (S-1) 0.82 1.9 0.18 0.070 0.038 KM (MM) 0.08 1.3 4.2 24.0 0.35 1 Kcat/ KM (mM-¹ s¯¹) 10.3 1.5 0.04 0.003 0.11 The value of kcat for N-Ac-Phe-OC2H5 is two-fold greater than that for the L-tryptophanyl analog and more than 10-fold greater than the value of kcat for the ester substrate…arrow_forwardThere are parts A-C for this picture included. A) What type of enzyme is Malate Dehydrogenase? choices: Hydrolase, Isomerase, Ligase, Oxideoreductase, Transferase, or Translocase B) Which of the following statements are true in biochemical standard conditions? There can be more than 1. Choices: The reaction is spontaneous since ∆G°' is positive The reaction is spontaneous since ∆G°' is negative The reaction is not spontaneous since ∆G°' is positive The reaction is not spontaneous since ∆G°' is positive The equilibrium favors products since K is greater than 1 The equilibrium favors reactants since K is greater than 1 The equilibrium favors products since K is less than 1 The equilibrium favors reactants since K is less than 1 The reaction is always at equilibrium C) If the concentration of Oxaloacetate is 10^7 times lower than the concentration of Malate D.,Is the reaction Spontanuous? Choices: No, because RTInQ is very positive Yes, because RTlnQ is very…arrow_forwardName, draw and describe the organic product of the reaction between 2-methylbut-1-ene and H2O in the presence of H2SO4 and provide a clear rationale as to why this is the major product of the reaction.arrow_forward
- 10. Procaine is known to undergo faster metabolic hydrolysis than procainamide. Give explanation. -NET2 -NEt2 HN H2N- H2N- Procainamide Procaine 11. In phase 2 metabolic transformation chloramphenicol the primary alcohol is converted to a glucuronide conjugate. Draw chemical structure of the product. Provide the name of enzyme that catalyze this metabolic reaction. он ? CHCI2 O2N- ноarrow_forwardIn many biochemical reactions which involves the formation of an enolate intermediate, the carbonyl oxygen of the substrate is coordinated to a divalent metal ion (usually zinc or magnesium) in the active site. Explain with structural drawings, how this ion-dipole interactions affect the acidity of the a-protons?arrow_forwardGive at least 5 examples of biological compounds having a ketone functional group and identify the biochemical importance of each compound.arrow_forward
- 4- Salicylamides are inhibitors for an enzyme called scytalone dehydratase. SAR shows that there are three important hydrogen bonding interactions. Explain whether you think quinazolines could act as a bioisostere for salicylamides. HBA OH O 00 N NHR Quinazolines Salicylamides Ph HBA 5- Structure IX (X = NH) is an inhibitor of a metalloenzyme called thermolysin and forms interactions as shown. Explain why the analogue (X = O) has reduced binding affinity by a factor of 1000 and why the analogue (X = SH) has roughly the same binding affinity. Structure IX H HBD Ala-113 Rarrow_forward5-phosphate) and an aldopentose (ribose-5-phosphate) to an aldotriose (glyceraldehyde-3-phosphate) and a ketoheptose (sedoheptulose-7-phosphate). Notice that the total number of carbons in the reactants andproducts is the same (5 + 5 = 3 + 7). Propose a mechanism for this reaction. xylulose-5-PCH2OH CH2OPO32−C OHOHHOH sedoheptulose-7-Parrow_forwardDraw the amino acid product that results from a transamination reaction involving pyruvate and glutamate, shown in the image below. Draw the product as it would occur at the pH of most body fluids, including all charges. You do not need to draw the other product of the reaction. O NH3 3D CH3-C-C-o-+ 0-CH-CH CH2-C-0--→? pyruvate glutamate Draw the molecule on the canvas by choosing buttons from the Tools (for bonds), Atoms, and Advanced Template toolbars. The single bond is active by default. Include all hydrogen atoms and charges. > View Available Hint(s) CONT. H. N. NV CI Br 1] -C- C1 SIarrow_forward
- 22) answer the following question.. Refer to the kinetic scheme for competitive inhibition and the structures shown below to E+S ES E +P -co- CO- 1 2 EI Compound 1 was determined to act as a competitive inhibitor through standard inhibition studies. Structural studies did not show any resemblance to the transition state. Compound 2 was also determined to act as a competitive inhibitor. Structural studies showed that it does resemble the transition-state. The K, constant is used to assess relative affinity of inhibitors for enzymes. That is, each compound has its own K, value. We can interpret K, the same way we do with Ka values. True or False: K, > Kµ2. Briefly explain your answer.arrow_forwardThe substitution of His 64 of carbonic anhydrase II with Ala results in a sharp decrease in the activity of the enzyme in HEPES buffer (molecular weight of HEPES = 238.3 g/mol). However, increasing concentrations of imidazole (molecular weight = 68.1 g/mol) restores the reaction rate close to that of the wild-type enzyme. Propose an explanation for these results.arrow_forward1. Please fully explain (use illustrate where appropriate) the Modes of Enzyme Catalysis exemplified by the serine protease: Chymotrypsin. In your answer discuss employing the illustration whenever possible: the overall reaction mechanism, stability of the reaction transition state, proximity and orientation effects, acid-base catalysis, and covalent catalysis. (c) (0) Ap Asp Toe His Asp 10 C-N bond cleavage HN Ho Ser Ger Binding of substi 196 Ser Gly alto video LBHB NH Sere HAR Proton donation by H (h) Fel of amino product yest OHN Hig Ser Ap (0) Formation of covalent (ES) Alp Me complex Seriosarrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





