
Concept explainers
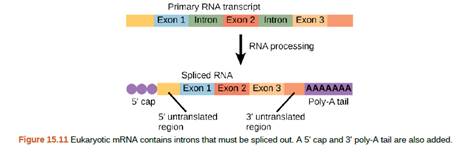
Figure 15.11 A scientist splices a eukaryotic promoter in front of a bacterial gene and inserts the gene in a bacterial chromosome. Would you expect the bacteria to transcribe the gene?
To analyze:
Whether the bacterial gene having eukaryotic promoter gets transcribed or not.
Introduction:
Eukaryotic promoters are different from prokaryotic promoters so the bacteria would not transcribe the gene.
Explanation of Solution
In case of transcription of genes, i.e. formation of RNA (mostly m-RNA) from DNA, promoters play an essential role. Promoters are the specific DNA sequences present near the starting point of transcription of a particular gene. Promoters help in the binding of RNA polymerase and transcription machinery to the specific point in the gene to initiate the process of transcription. They are mainly located upstream of a gene, towards 5’ end of the anti-sense strand. As these are specific recognition sequences, eukaryotic promoters are different from prokaryotic promoters.
Prokaryotic promoters have three essential elements:-10 elements,-35 element and UP (upstream promoter) element.
Eukaryotic promoters mainly contain a TATA box of six nucleotides (consensus sequence is 5’ TATAAA 3’) located around-30 (with reference to initiation site of transcription +1), while prokaryotic promoters have its counterpart-10 elements or Pribnow box of six nucleotides (consensus sequence is 5’ TATAAT 3’) located at-10. Both these have A-T rich elements. Prokaryotes also have one more promoter sequence of six nucleotides at-35 which is (5’ TTGACA 3’) called as-35 elements. Pribnow box or-10 element is essential in prokaryotes for the initiation of transcription and-35 element helps in increasing the transcription rate.UP element is A-T rich sequence located between-40 to-60.
Other conserved sequences in eukaryotic promoters are CAAT box (GGCCAATCT) around-80 and GC-rich boxes (GGCG), and octamer boxes (ATTTGCAT) located further upstream.Basically, eukaryotic promoters are much more complex and larger in structure than the prokaryotic promoters.Sigma factor (s factor) of prokaryotic RNA polymerase holoenzyme recognizes and binds to the promoter interaction site to initiate transcription.
So, a bacterial gene having eukaryotic promoter would not transcribe because eukaryotic promoter has different recognition sites/consensus sequences than the required prokaryotic recognition sites. RNA polymerase, transcription factors and other proteins of host bacterial chromosome would not be able to identify the eukaryotic promoter sequences and transcription of the gene would not occur.
So, the bacteria would not transcribe the gene with the eukaryotic promoter as they are different from the prokaryotic promoters.
Want to see more full solutions like this?
Chapter 15 Solutions
Biology 2e
Additional Science Textbook Solutions
College Physics
Concepts of Genetics (12th Edition)
Campbell Essential Biology with Physiology (6th Edition)
Fundamentals of Anatomy & Physiology (11th Edition)
Anatomy & Physiology
- Put the following processes in order of their occurrence during expression of a eukaryotic gene: a. mRNA processing c. transcription b. translation d. RNA leaves nucleusarrow_forwardConsider the generalized structure of a eukaryotic gene, and the RNA it encodes. Of the options listed below, which one set of features determines the length of the primary RNA in base pairs, before any processing has occurred? The exon and intron sequences The promoter and enhancer sequences The start and stop codons for translation The transcription start and termination sequencesarrow_forwardHere is a schematic map with a scale of a eukaryotic gene. How long is the processed mRNA transcript produced from this gene? Termination Start codon Stop codon sequence -35 -10 Exon 1 Intron 1 Exon 2 Intron 2 Exon 3 1 2 3 4. Genomic sequence (kb) (k = kilo = 1,000) 2200 bases 3200 bases 2000 bases 4200 basesarrow_forward
- Refer to the double stranded DNA molecule with the sequence below to answer the following questions: 5’ATATGGGTCTCGATAGGGCTGTTTTCTCCGGC 3’ 3’TATACCCAGAGCTATCCCGACAAAAGAGGCCG 5’ Which strand functions as the transcription template, the top one or the bottom one? Explain your reasoning. What is the mRNA transcript and polypeptide from this strand? In the space below, copy the DNA strand that is transcribed, and write the mRNA transcript and polypeptide chain below it. Align the mRNA and polypeptide so that it is clear which DNA bases they came from. DNA strand: mRNA: amino acid sequence:arrow_forwardSeveral different nucleic acids are involved in the process of getting a protein produced from a gene. DNA contains the "genetic code" for the protein. DNA is double-stranded, but only one strand is transcribed into MRNA. The MRNA then goes into the cytoplasm where it is translated into protein with the help of TRNA. At each stage of the process, there is base complementarity (A pairs with T/U and C pairs with G) between the nucleic acids involved to ensure the integrity of the DNA blueprint for the protein being produced. Therefore, some of the four strands of nucleic acids involved will match (except U replaces T in RNA) and some will have base complementarity. Indicate whether there is matching (1) or base complementarity (2) between the following nucleic acids. DNA sense strand and MRNA DNA sense strand and tRNA DNA antisense strand and MRNA MRNA and TRNAarrow_forwardWhich one of the following features could you use to determine the length of the RNA that is produced based on the gene, before it is processed? The exon and intron sequences The promoter and enhancer sequences The start and stop codons for translation The transcription start and termination sequencesarrow_forward
- The sequence below shows a mRNA sequence derived from a template strand of a DNA molecule: DNA sequence 1: CTTTTTTGCCAT DNA sequence 2: ACATCAATAACT DNA sequence 3: TACAAGGGTTCT Determine the mRNA sequence that you would derive from transcription of each strand Using the genetic code table, show the amino acid sequence that will be encoded by these mRNAs For the 3 DNA sequence, what will be the sequence of the complementary base after the process of replication?arrow_forwardUsing the example above, transcribe the following DNA strand into mRNA and translate that strand into a polypeptide chain, identifying the codons, anticodons, and amino acid sequence.arrow_forwardGiven the following DNA sequence of the template (i.e. noncoding) strand for a given gene: 5'ATTGGCTGTTAGAGCGGCCGTCTAAACATCGTTGGA3' Part A) Write the mRNA that will be transcribed from the DNA sequence above (be sure to label the 5' and 3' ends) Part B) Use the genetic code to write the peptide sequence translated in a cell from the mRNA synthesized in part A. Please use the 3 letter abbreviation for each amino acid.arrow_forward
- You obtained the sequence of the frog gene X you amplified in Question #16 through a process called automated sequencing. In automated sequencing, you are given a printout of the sense strand of your DNA. The printout is shown below. The first thing you need to do is use the correct reading frame. Having done this, the next thing to do is to write out the mRNA sequence using this sense strand reading frame. The last thing to do is to translate the sequence. Do these steps in the space below. The reading frame DNA sequence is: 2.The mRNA sequence is: The polypeptide sequence is:arrow_forwardYou are studying the rate of transcription of a particular eukaryotic gene. When the DNA located several thousand bases upstream from the gene is removed, the transcription rate of the gene decreases dramatically. How would you interpret these results?arrow_forwardCompare the composition of genes and upstream regions of DNA in bacteria, Archaea and eukaryotes in table formatarrow_forward
 Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax

