
Concept explainers
Hershey–Chase Experiments The graph shown in FIGURE 8.5 is reproduced from an original 1952 publication by Hershey and Chase. Bacteriophage were labeled with radioactive tracers and allowed 10 infect bacteria. The virus–bacteria mixtures were then whirled in a blender to dislodge any viral components attached to the exterior of the bacteria. Afterward, radioactivity from the tracers was measured.
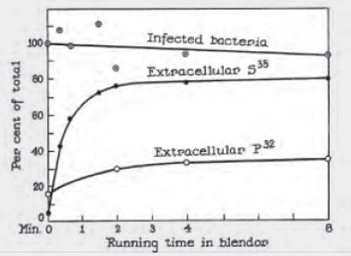
FIGURE 8.5 Detail of Alfred Hershey and Martha Chase’s 1952 publication describing their experiments with bacteriophage.
“Infected bacteria” refers to the percentage of bacteria that survived the blender.
Before blending what percentage of each isotope. 35S and 32P, was extracellular (outside the bacteria)?
To explain: The percentages of
Concept introduction: DNA is the organic molecule that carries information from one generation to another. Several experiments were to be conducted to prove that DNA is the genetic material. One such experiment is the Hershey–Chase experiment commonly known as the blender experiment. Bacteriophages are viruses that infect the bacteria and take up their translation machinery for their life cycle. This property was exploited to prove that DNA is a genetic material along with the help of radioactive isotopes.
Explanation of Solution
A graph was plotted with running time in the blender as X-axis and percentage of radioisotopes and infected bacteria in the Y-axis. Three scatter plots were drawn one for each radioactive tracer
Before blending the mixture that is, at 0 minute the percentage of radioactive tracers were measured. The percentages of
Want to see more full solutions like this?
Chapter 8 Solutions
Biology: The Unity and Diversity of Life (MindTap Course List)
Additional Science Textbook Solutions
Microbiology with Diseases by Body System (5th Edition)
Human Anatomy & Physiology
Microbiology Fundamentals: A Clinical Approach
Anatomy & Physiology: The Unity of Form and Function
SEELEY'S ANATOMY+PHYSIOLOGY
Prescott's Microbiology
- HersheyChase Experiments The graph shown in FIGURE 8.5 is reproduced from an original 1952 publication by Hershey and Chase. Bacteriophage were labeled with radioactive tracers and allowed 10 infect bacteria. The virusbacteria mixtures were then whirled in a blender to dislodge any viral components attached to the exterior of the bacteria. Afterward, radioactivity from the tracers was measured. FIGURE 8.5 Detail of Alfred Hershey and Martha Chases 1952 publication describing their experiments with bacteriophage. Infected bacteria refers to the percentage of bacteria that survived the blender. After 4 minutes in the blender, what percentage of each isotope was extracellular?arrow_forwardHersheyChase Experiments The graph shown in FIGURE 8.5 is reproduced from an original 1952 publication by Hershey and Chase. Bacteriophage were labeled with radioactive tracers and allowed 10 infect bacteria. The virusbacteria mixtures were then whirled in a blender to dislodge any viral components attached to the exterior of the bacteria. Afterward, radioactivity from the tracers was measured. FIGURE 8.5 Detail of Alfred Hershey and Martha Chases 1952 publication describing their experiments with bacteriophage. Infected bacteria refers to the percentage of bacteria that survived the blender. The extracellular concentration of which isotope increased the most with blending?arrow_forwardHersheyChase Experiments The graph shown in FIGURE 8.5 is reproduced from an original 1952 publication by Hershey and Chase. Bacteriophage were labeled with radioactive tracers and allowed 10 infect bacteria. The virusbacteria mixtures were then whirled in a blender to dislodge any viral components attached to the exterior of the bacteria. Afterward, radioactivity from the tracers was measured. FIGURE 8.5 Detail of Alfred Hershey and Martha Chases 1952 publication describing their experiments with bacteriophage. Infected bacteria refers to the percentage of bacteria that survived the blender. Do these results imply that viruses inject DNA or protein into bacteria? Why or why not?arrow_forward
- Procedure: 1. Using the DNA provided transcribe DNA into mRNA. 2. Use the mRNA strand you created and break it up into codons. 3. Plug the codons into the amino acid chart to determine the correct amino acid needed to build that protein. 4. Identify the protein you made by comparing the sequence to the pictures 5. Answer the questions for each protein molecule you build before moving on to the next. Protein 1: DNA A AGACCGTATAC mRNA Amino Acid Sequence 1. Which kind of protein molecule did this gene make? 2 How does this protein help the body maintain homeostasis2arrow_forwardTRY TO KEEP IN SHORT AND USE OWN WORD FOR THIS QUESTION You are studying a type of bacteria isolated from the acidic water runoff of a mining operation. You subject two batches of the same bacteria type to different environmental growth conditions. One batch is grown at pH 2, while the other is grown at pH 7. All other environmental parameters are kept identical between the two batches. You then collect their proteins and run a Western blot using an antibody that binds to a proton efflux pump protein (which actively expends energy to pump protons out of a cell). How would you characterize the information obtained in this experiment? What does it tell you, and why is that potentially valuable information?arrow_forwardWhat were the findings of Hershey and Chase's experiments with bacteriophages? Select all that apply. Radiolabeled viral protein was found only inside of bacteria cells. Radiolabeled viral protein was found only outside of bacteria cells. Radiolabeled viral DNA was found only outside of bacteria cells. Radiolabeled viral DNA was found only inside of bacteria cells.arrow_forward
- Imagine that you are a student in Alfred Hershey and Martha Chase’s lab in the late 1940s. You are given five test tubes containing E. coli bacteria infected with T2 bacteriophages that have been labeled with either 32P or 35S. Unfortunately, you forget to mark the tubes and are now uncertain about which were labeled with 32P and which with 35S. You place the contents of each tube in a blender and turn it on for a few seconds to shear off the phage protein coats. You then centrifuge the contents to separate the protein coats and the cells. You check for the presence of radioactivity and obtain the following results. Which tubes contained E. coli infected with 32P-labeled phage? Explain your answer. Tube number Radioactivity present in 1 Cells 2 Protein coats 3 Protein coats 4 Cells 5 Cellsarrow_forwardquestion: Can you summarize and explain for me what you want to tell in the article below? When I read it myself, I do not understand exactly what is meant by the article. It would be nice if you could highlight the important points. You can use them in a figure or diagram to explain. thank you and hava a nice day :) Article: Biomimetic Engineering of Nanodelivery Systems: Artificial Viruses in the Making In an effort to engineer the next generation of nanoscale vectors, scientists have moved from using inorganic components aimed at obtaining inert structures to utilizing biological building blocks that are able to convey additional functionalities to the resulting construct. To cope with the complexity of the body and to evade the multiple layers of defense that tissues and organs have, it is critical to rely on the ability of certain materials to interact with, rather than to eschew, the biology of our body. Every NP system conceived to date faces one common fate: whether injected,…arrow_forwardDNA Extraction by Alkaline Lysis Procedure: 1. Spin 1.5 ml of cells in a microcentrifuge at maximum speed (12,000 rpm) for 20s to pellet. Remove the supernatant completely with a Pasteur pipet or a plastic pipettor tip. The spins can be performed at 4C or at room temperature. Longer spins make it difficult to resuspend cells. 2. Resuspend pellet in 100µl GTE solution and let sit 5 min at room temperature. Be sure cells are completely resuspended. 3. Add 200µl NaOH/SDS solution, mix by tapping tube with finger, and place on ice for 5 min. 4. Add 150µl potassium acetate solution and vortex at maximum speed for 2s to mix. Place on ice for 5-15 min. Be sure mixing is complete. 5. Spin 3 min at 12,000 rpm to pellet cell debris and chromosomal DNA. 6. Transfer 0.4 ml supernatant to a fresh tube, mix it with 0.8 ml of 95% ethanol or 0.4 ml isopropanol, and let sit 2 min at room temperature precipitate nucleic acids. 7. Spin at 12,000rpm for 3 min at room temperature to pellet plasmid DNA and…arrow_forward
- A media contains 2 E. coli cells which have a generation time and a replication time of 60 and 40 mins respectively. How many cells will be produced after 5 hours?arrow_forwardPipetting 1. Explain why a micropipettor is an important instrument in biochemistry labs. 2. Describe the basic components (and function) of a micropipettor. Bacterial Techniques 1. Define the following. (a) Serial Dilution (b) Streak Plating (c) Spread Plating Transformation 1. Define bacterial transformation and explain why it is an important method in biochemistry labs. 2. Describe (figure, narrative) a plasmid and describe the basic components of a plasmid. 3. What role does CaCl2 play in bacterial transformation? 4. What role does heat shock play in bacterial transformation? Plasmid Isolation 1. Describe the roles the following play in plasmid purification. (a) Lysis Buffer (b) Neutralization Buffer (c) Elution Bufferarrow_forwardquestion: Can you summarize and explain for me what you want to tell in the article below? Can you explain the figure? When I read it myself, I do not understand exactly what is meant by the article. It would be nice if you could highlight the important points. You can use them in a figure or diagram to explain. thank you and hava a nice day :) Article: Nanotechnology Tools to Detect SARS-CoV-2 Standard procedures for detecting the virus from nasopharyngeal and/or oropharyngeal swabs have been reviewed recently and are primarily based on reverse transcription polymerase chain reaction (RT-PCR). Here, we would like to mention some preliminary ideas on nanotechnology-based assays to monitor the presence of SARS-CoV-2. A simplified test and variants thereof to detect viral proteins (e.g., HIV or influenza virus) without the need for expensive equipment is based on the color change of Au NPs bound to antibodies. Similar to the enzyme-linked immunosorbent assay (ELISA) antibodies coupled…arrow_forward
 Biology: The Unity and Diversity of Life (MindTap...BiologyISBN:9781305073951Author:Cecie Starr, Ralph Taggart, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology: The Unity and Diversity of Life (MindTap...BiologyISBN:9781305073951Author:Cecie Starr, Ralph Taggart, Christine Evers, Lisa StarrPublisher:Cengage Learning
